
Review Article
Volume-5 Issue-1, 2026
Honouring a Scientific Pioneer: The Enduring Legacy of Professor Soon Guan Tan (1947–2023) in Malaysian Genetics, Biodiversity, and Environmental Science
Received Date: December 20, 2025
Accepted Date: January 04, 2026
Published Date: January 08, 2026
Journal Information
Abstract
We are privileged to honour Professor Soon Guan Tan (Prof SGT), who’s pioneering genetics and biodiversity studies in Malaysia laid the groundwork for subsequent research. The comprehensive analysis of publications, theme clusters, and research evolution from 1976 to 2021 shows how his work shaped national scientific capacity and changed biological research in numerous disciplines. Prof SGT established Malaysian human and animal genetic baselines using biochemical and population genetic methodologies early in his career. He promoted molecular biological approaches to modernize Malaysian laboratories and enhance conservation genetics, wildlife genomics, agricultural, and environmental monitoring studies. His latter research integrated genetics with ecosystem-level thinking to improve environmental stewardship and resource management. More than his scientific output, his mentorship, leadership, and intellectual generosity have shaped generations of scientists. This paper honours his legacy and recognizes his career effort as the foundation for Malaysian genetic and ecological research.
Key words
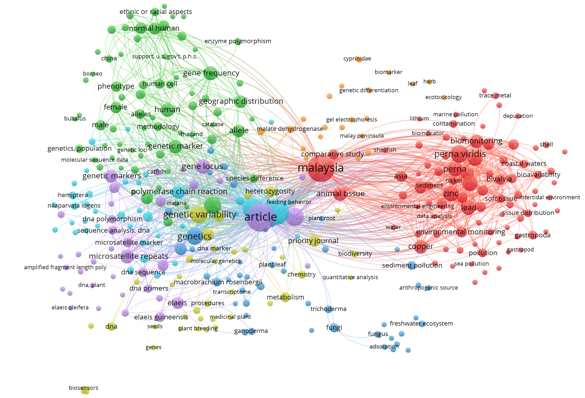 |
| Figure 1: Keyword co-occurrence network of Professor Soon Guan Tan’s publications (1976–2021) generated using VOSviewer, showing major thematic clusters across genetics, molecular markers, biodiversity, and Malaysian population studies. |
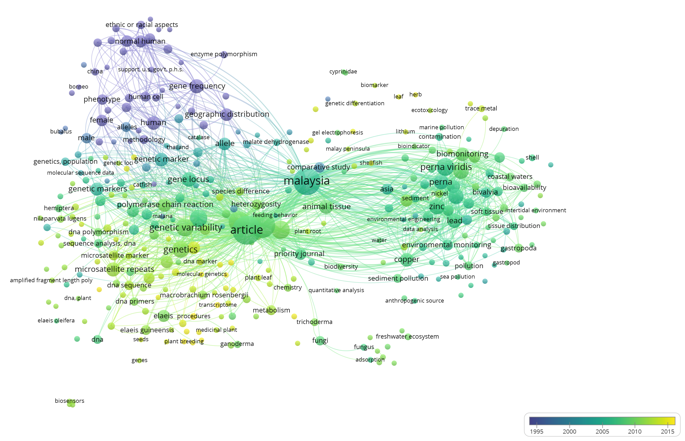 |
| Figure 2: Temporal overlay visualisation of keyword co-occurrence in Professor Soon Guan Tan’s publications (1976–2021), illustrating the chronological evolution of research themes from early genetic marker studies to later biodiversity and ecotoxicology applications. |
Introduction
The scientific landscape in Malaysia in the 1970s was far different. Colleges in Southeast Asia were then just beginning to introduce population genetics and modern molecular biology. This climate helped Professor Soon Guan Tan (Prof SGT) pioneer investigations on biochemical and genetic marker studies in Malaysian research. His first single-authored article, Human saliva esterases: Genetic investigations (Tan, 1976), launched a distinguished scientific career and opened a new avenue of research for the country. This pioneering work demonstrated Malaysian salivary esterase genetic variability and, importantly, one of the first empirical frameworks for population structure based on biochemical genetics.
Prof SGT's expanding research on polymorphic markers in saliva, blood, and other tissues revealed the effect of this early work. Thus, late 1970s collaborative research of acid phosphatases, amylase polymorphisms, and mitochondrial enzyme markers in Malays, Chinese, Indians, Aboriginal communities, and Sabah ethnic groups created the country's first genetically informed map of human variety. These comparative analyses found population genetics mapping. Previously rare in the region, these pioneering efforts established Malaysia as a population genetic study center. His methodological innovation in using markers showed how genetic evidence underlined population history, migration, and variation.
When new technologies started pouring in from around the world in the 1990s, Prof SGT easily shifted his study into protein electrophoresis, PCR-based markers, microsatellites, and transcriptomics. Later works on Malaysian mahseer microsatellite characterisation (Chew et al. 2021), primate transcriptome genic-SSR finding (Chang et al. 2019), and wild Macaca fascicularis RNA sequencing (Ee-Uli et al. 2018) had shown how he applied advanced genomic methods. These investigations moved Malaysian genetics beyond population-level marker studies into wildlife genomics, aquaculture genetics, and molecular diagnostics. His leadership in Malaysia ensured that Malaysian laboratories kept up with international scientific development in high-resolution molecular technologies.
In addition to human and wildlife genetics, Prof SGT advanced biodiversity conservation, agricultural genetics, environmental monitoring, and aquatic ecotoxicology. His molecular tools were used to study oil palm and durian, medicinal plants, and animal disease and environmental investigations. Biomonitoring and environmental governance benefited from his research on Diadema setosum developmental sensitivity and other marine creature environmental reactions. These publications indicate the remarkable range of a scientist who commenced analyzing salivary esterases in 1976 and diversified into a multi-disciplinary portfolio shaping Malaysian research.
This paper examines Prof. SGT's contributions to scientific scholarship in order to determine how they have changed over three decades with respect to conceptual, methodological, and disciplinary dimensions. In particular, an examination of his publication record, thematic clusters of research activity, and the impact of his research across disciplines, provides evidence of how Prof. SGT has advanced population genetics in Malaysia, strengthened national capacity in molecular biology, established a framework for research on biodiversity and conservation, and expanded his research into environmental monitoring, ecotoxicology, and genomic sciences. Through this examination, we hope to demonstrate the influence of Prof. SGT's legacy on the future of Malaysian biologists, geneticists, and environmental scientists as well as the enduring relevance of his views and methodologies in science.
Methodology
The scientific legacy of Prof. SGT was examined through the analysis of bibliometric data and historical records, as well as qualitative techniques used to assess the conceptual and disciplinary aspects of his work. The methodological component involved two phases: (1) retrieving bibliographic (publishing and citation) records using the Scopus database (provided by Elsevier) for the past two decades, and (2) investigating the conceptual and disciplinary significance of his work qualitatively. Despite the advantages of using Scopus for bibliometric consistency, standardized metadata, and reliable author disambiguation, reliance on a single database represents a methodological limitation. Scopus does not comprehensively index all local, regional, or non-English language journals, particularly those publishing context specific or policy oriented research. As a result, some locally grounded studies may not be captured in the dataset, potentially introducing coverage bias. This limitation is acknowledged, and the findings should therefore be interpreted as representing patterns within the Scopus indexed literature rather than the entirety of global scholarship on the topic.
A bibliometric search was carried out in Elsevier's Scopus database on 11 December 2025. The author’s name searched: "Tan" and "Soon Guan." In order to ensure comprehensiveness of the search results and author disambiguation, the results had been cross-validated with the author's Scopus profile: Scopus ID: 7403366474; ORCID: 0000-0001-9960-4493. The search resulted in 248 research articles, book chapters, and conference papers that were all indexed from 1976 to 2021. There were 5,453 citations at retrieval from 4,047 citing papers, with an h-index of 39. These metrics quantitatively supported the profiling of Prof SGT’s scientific influence.
In order to avoid misattributions of publications by Prof SGT, each record was compared and cross-checked against author affiliations, co-author networks, research themes, and institutional identifiers. The name "Tan" is fairly common in Malaysia and Singapore, manual verification was therefore use to prevent erroneous merging of the author profiles or misclassification of publications. Only publications consistently linked to Universiti Putra Malaysia or to confirmed collaborators were retained. Before analysis, duplicates, and mis-indexed entries were removed.
For Prof SGT, the bibliometric measures included publication count, citations, and citations per document, H-index, and publication distribution by year. These variables enabled the modeling of his career duration, growth in research productivity, and the impact of important papers. Since Prof SGT's contributions span genetics, molecular biology, biodiversity, population studies, and agricultural sciences, his citation numbers were not field-normalized.
Keyword co-occurrence analysis was conducted using VOSviewer (version 1.6.20), a well-known bibliometric tool for building and visualizing networks (Van Eck & Waltman, 2010, 2014, 2017), to determine the topic structure of Prof SGT's research. Using the entire dataset, 1,606 author and indexed keywords were retrieved. Threshold study indicated that 344 keywords had sufficient occurrences for reliable mapping. Through clustering keywords into thematic groupings, some key areas of study emerged: population genetics, electrophoretic markers, Southeast Asian biodiversity patterns, agricultural genetics, and conservation biology. The shifting scholarly focus and contributions of Prof SGT to genetic techniques in Malaysian biological sciences were traced.
The qualitative investigation into Prof SGT's career trajectory from the 1970s to the early 2020s involved a review of landmark papers, an examination of collaborative networks, and the positioning of his scientific contributions within the broader development of genetics and biodiversity research in Malaysia and Southeast Asia. In this study, the author established selection criteria that evaluate selected papers based on methodological innovation, conceptual contribution, and influence on research within molecular genetics, population biology and conservation genetics. Citation analysis was used to contextualize bibliometric data by ensuring they were interpreted within their disciplinary context, rather than relying solely on publications and citations.
Quantitative and qualitative results were brought together to create the narrative of Prof SGT's science: what he contributed to genetics early in Malaysia, who were the scientists he trained on molecular genetics, and what role he played in fostering cross-disciplinary collaborations. A consideration of citation trajectories and modern relevance was also included when synthesizing the data.
Results
VOS viewer analysis identified seven clusters covering the period from 1976 to 2021, which identified the intellectual structure of the scientific legacy of Prof SGT. These clusters, as presented in different colors in the network visualization of Figure 1 and temporally enriched in Figure 2, extend his contributions from population genetics to biodiversity studies into ecological and environmental applications. These clusters represent thematic breadth, though methodological continuity was observed based on genetic markers and population-based analysis.
Green Cluster 1: Fundamentals of Genetics and Gene Frequency Studies
This cluster consists of early to mid-career keywords: gene frequency, genetic diversity, heterozygosity, markers, allele distribution, and genotype. As Prof SGT conducted population genetics and evolutionary mechanisms research in Malaysian fauna since the late 1970s to the 1990s, there are a lot of activities in the temporal overlay. Electrophoresis, isozymes, and microsatellite markers indicate his methodological leadership in introducing molecular genetic technologies to Malaysian biology. Network richness indicates that this cluster reflects his pioneering research on genetics across diverse taxa. Chronological change shows that these would be core studies to support ecological and conservation research.
Blue Cluster 2: Molecular Methods and Marker Development
This cluster is Prof SGT’s study methodology, represented by such terminology as PCR, DNA polymorphism, RAPD, AFLP, microsatellite DNA, and sequencing. Electrophoresis was first developed in the 1980s, while the PCR-based approaches took off until the late 1990s and early 2000s (Figure 2). These keywords reflect the classical genetic markers shifting to modern molecular approaches, indicating that Prof SGT quickly adopted and applied the new technologies in research. He focused on the arker development for species identification, genetic variability evaluation, and phylogenetic reconstruction in this cluster. The high connectedness between this cluster and Cluster 1 shows the methodological continuation of his genetic research.
Purple Cluster 3: Agriculture and Crop Genetics
Plant breeding, crop genetics, microsatellites, rice, maize, genetic improvement, and agro-biodiversity are included. This minor cluster of Prof SGT 's contributions in agricultural genetics contrasts with large animal-oriented clusters. Activities appeared from 2000 to 2010, reflect Malaysia's research priorities on crop development, food security, and genetic resource conservation. Network connectivity indicates the transfer of methodologies from animal molecular markers to plant systems. This cluster proves that Prof SGT was a geneticist who contributed to building Malaysia's agricultural genetics for scientific modernization.
Light Blue Cluster 4: Malaysian Biodiversity and Species Distribution
Prof SGT generated knowledge on regional biodiversity through the Malaysia-centered cluster, species distribution, population structure, comparative studies, animal tissues, and morphology. Temporal evidence points to this cluster's mid-1990s origin and growth in the 2010s. Comparative genetic surveys were carried out in terrestrial and aquatic species. His works quickly became one of the standards to characterize biodiversity in Malaysia, especially those economically or ecologically important species. Strong bridge links of this cluster to Clusters 1 and 2 suggest that genetic technologies were utilized to investigate biodiversity concerns, hence locating Prof SGT at the crossroads between genetics and conservation biology.
Red Cluster 5: Aquatic Biology, Perna viridis, and Coastal Ecosystems
The most distinctive cluster in Figure 1 includes the green-lipped mussel Perna viridis, biomonitoring, ecotoxicology, sediment, pollution, bioaccumulation, and coastal ecology. This cluster starts in the early 2000s and it represents Malaysia's interest in marine pollution, aquaculture, and coastal protection. Prof SGT pioneered the use of genetic and biochemical methods to aquatic species, especially mussels, becoming one of Malaysia's subjects of biomonitoring research. The characteristics of density and cohesiveness of this cluster imply a highly developed multi-disciplinary cooperation for an established program involving genetics, environmental chemistry, ecotoxicology, and marine ecology.
Yellow Cluster 6: Applications to Environmental Monitoring and Ecotoxicology
Within this cluster, the components include heavy metals, trace elements, sediment quality, anthropogenic input, contamination, and ecological risk. Although there is a connection with the mussel-centric cluster, this cluster appears to represent a broader scope of environmental interest. Visibility of this cluster, as reflected in Figure 2, increases from late 2000s to 2010s. Prof SGT's transition into applied environmental science and ecosystem health assessment is reflected here. The scope and analytics involved in this study in coastal management and environmental policy gave rise to terms such as risk assessment and quantitative analysis. Prof SGT contributed to bringing genetic knowledge into practical monitoring mechanisms.
Orange Cluster 7: Ecological Processes, Trophic Interactions, and Ecosystem-Level
Studies that were small in scope but but significant to the conceptual framing are ecosystem, trophic linkages, feeding behaviour, bioindicators, and ecological processes. Temporal mapping indicates moderate but constant activity since the 1990s. While not his main focus, this cluster reveals how Prof SGT's genetic and biomonitoring studies have framed ecological interpretations. The cluster's visibility within the network supports the interdisciplinary portfolio presented by Prof SGT, illustrating his influence extends outside molecular genetics into ecosystem-level understanding. Location at the boundary of the genetics and ecotoxicology clusters illustrates how this cluster integrates organismal biology with environmental processes.
Discussion
Laying the Foundations of Malaysian Population Genetics
The earliest clusters indicate that the studies of allele frequencies, heterozygosity, and genetic markers by Prof SGT had laid the foundations for Malaysian molecular biology. His early studies on biochemical polymorphisms in the saliva, blood, and other tissues of Malaysian ethnic groups demonstrated that enzyme and protein markers can be used to characterize population structure and variability. These soon expanded into detailed surveys of genetic markers such as glutamic oxaloacetic transaminase, amylase, phosphoglucomutase, esterases, and superoxide dismutase across Malay, Chinese, Indian, Aboriginal, Kadazan, and other communities, providing one of the first systematic portraits of genetic diversity in the country. When genetic research in Southeast Asia was still in its infancy in the 1970s and early 1980s, his innovative use of electrophoresis and isozyme markers established the methodological tools and conceptual frameworks necessary to study natural populations.
Studies on cerumen polymorphism, salivary peroxidase, and Gc subtypes also expanded the comparative perspective and relationships of population and gene flow in Malaysia and Indonesia. His early publications established the country's first coherent body of knowledge on evolutionary patterns across Malaysian fauna and human populations, laying the groundwork for later genetic and ecological research in Malaysia.
Novel Molecular Techniques and Research Capacity
As a Professor, SGT's adaptability and innovation were overt as the change came in during the 1990s, when molecular biology took over the research field globally. This shift from electrophoretic markers towards PCR, microsatellite, DNA polymorphism, and high-throughput sequencing demonstrates the adaptability of Malaysian science to keeping pace with global breakthroughs.
His later work on microsatellite characterization for Malaysian mahseer broodstock management best illustrated the use of highly informative markers within fisheries genetics and aquaculture. He developed and applied SSR and genic-SSR markers, QTL mapping, and genetic diversity assessments in oil palm, durian, and Andrographis paniculata, underpinning molecular-assisted selection and breeding platforms. Refining multiplex PCR assays for the discrimination of closely related Cymbopogon species and applying inter-simple sequence repeat and chloroplast DNA markers in horticulture crops demonstrate his technical competencies, which were transferable across taxa. Malaysian laboratories entered the worldwide molecular genetics landscape when novel genic-SSR markers and molecular sex-identification techniques for painted storks and non-human primates were developed. The highly connected blue and purple clusters reflect a legacy of modernization whereby many postgraduate students have been trained in advanced methods of molecular biology and went on to lead laboratories and departments, enabling him to extend his influence through academic lineage and institutional capacity-building.
Enhance National Biodiversity Conservation and Comprehension of Science
The clusters of biodiversity, species distribution, and comparative studies serve to support the impacts of Prof SGT's research on ecological interpretation and conservation policy. Multi-species genetic and genomic studies of the wild populations, especially non-human primates such as Macaca fascicularis, provide valuable data on genetic diversity, population structure, and local adaptation in Peninsular Malaysia and Borneo. This work is complemented by research on marine invertebrates, for example, quantifying the effects of temperature and salinity on embryonic and early larval development in the black sea urchin Diadema setosum, to elucidate environmental limits to recruitment and resilience. Durian, roselle, and medicinal plant research pushed biodiversity genetics into agro-ecosystems and ethnobotanically significant species, embedding conservation in agricultural sustainability. Such research provided Malaysia for its first large-scale genetically informed view on biodiversity patterns across wild and farmed taxa, forming a critical basis for species-level conservation planning, protected area management, and protection. He linked molecular genetics to ecology and resource management; he can thus be regarded as a key architect of Malaysia's genetic biodiversity knowledge base.
Improvement of Environmental Monitoring and Mussel-Based Ecotoxicology
Prof SGT has developed his most meaningful legacy as an applied developmental biologist through a combination of genetic and biochemical aquatic monitoring of marine communities through Perna viridis and other coastal species combined with environmental monitoring. Environmental assessment encompassed plants and echinoderms as indicators for multi-taxa ecosystem health; however, molluscan biomonitoring remained as Prof SGT's main research focus. Temperature and salinity effects on Diadema setosum yielded sensitive developmental endpoints for the detection of tropical coastal water environmental change (Sarifudin et al., 2016; 2017). Trace element profiling in edible plants can inform human nutrition and soil quality assessment. Centella asiatica’s magnesium concentrations and habitat soils were studied in terrestrial contexts (Ong et al., 2017)
The synergistic cytotoxicity of cisplatin and bromelain in breast cancer cells demonstrated how biochemical and genetic tools could be applied in pharmacological and toxicological evaluation frameworks. Other projects also applied the biomonitoring concepts to aquaculture disease management and aquatic virology, such as Macrobrachium rosenbergii nodavirus virus-like particles and host response molecular characterization. Prof SGT integrated genetic expertise with environmental pollutants, developmental responses, and disease endpoints for pollution source allocation, ecosystem risk assessment, and environmental governance of Malaysia's coasts and agricultural landscapes.
Integrative Genetics, Ecosystem Thinking, and Scientific Stewardship Shape the Future
Ecological process and ecosystem-level keywords reflect Prof SGT's foresightedness. His later work employed high-throughput and omics-based approaches to integrate genetic data into ecological questions such as trophic relationship, habitat quality, and anthropogenic stresses. The lymphoid organs, kidney, and liver transcriptomic studies in wild cynomolgus macaques provided genomic resources for a very important biomedical and ecological model species, which would thus enable future research on disease ecology, environmental stress responses, and comparative genomics in Southeast Asia. The development of genic-SSRs from primate transcriptomes and the application of RNA-Seq in wildlife and agriculture would bridge conservation genetics, evolutionary biology, and functional genomics (Chang et al., 2019; Ee-Uli, 2018).
Advanced molecular improvement platforms for crops and medicinal plants, such as QTL mapping in oil palm and genoproteomics-assisted improvement in Andrographis paniculata, point to a future wherein genetic and ecological knowledge will undergird food security, pharmaceutical development, and ecosystem resilience. Multiplex PCR and DNA barcoding-style species diagnosis in Cymbopogon and other taxa reveal the increasing importance of rapid molecular diagnosis in biodiversity monitoring and biosecurity. From a scientific perspective, this involvement shows a way of developing a direct link between the scientific areas of genomics, environmental monitoring, and ecosystem modelling, providing direction for future research on climate change, habitat loss, and coastal degradation. Prof SGT developed a flexible system for studying population genetics, molecular technology, biodiversity science and environmental health that enabled Malaysian researchers to innovate and work across disciplines, maximizing benefits for biodiversity, society and the planet.
Conclusions
It is difficult to envision any younger professional matching the contributions made by Prof SGT in the study of genetics, biodiversity and environmental sciences in Malaysia. His findings are representative of the local life sciences, which he fostered from within through the development of human capital and scientific expertise. By establishing population genetics, Prof SGT involved in the search to establish the biochemical and genetic fingerprints that defined Malaysian humans and animals. His initial work laid the groundwork for future scientists in these areas. He embraced PCR and microsatellite markers, development of transcriptomics before the establishment of laboratories in the same region in Malaysia. He was also involved in establishing infrastructures to support molecular genetics, wildlife genomics and agriculture development. He innovated methodologies and shaped national biodiversity. As such, his multi-taxa studies within a conservation genetics framework advanced with an understanding of population structure, evolutionary relationships, and ecological robustness. Similarly, he employed mussel-based biomonitoring models and other ecological approaches to apply genetics to the management of aquaculture, assess the health of coasts, and exercise ecosystem stewardship. His career path inspires scientists. He described how to handle complex challenges in climate change, habitat degradation, and resource sustainability using genetics, ecology, molecular biology, and practical environmental science. He mentored, inspired, and empowered generations of students, collaborators, and researchers and left behind so much in terms of scholarship. Prof SGT leaves behind a body of work with scientific depth, disciplinary significance, and national relevance. His work will continue to shape Malaysian science, conservation, genetics, and environmental studies for decades to come.
Funding
This research received no external funding.
Institutional Review Board Statement
Not applicable.
Informed Consent Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflicts of interest.
References
- Chang W, Ee-Uli J, Ng WL, Rovie-Ryan JJ, Tan SG, Yong CSY (2019) Discovery of novel genic-SSR markers from transcriptome dataset of an important non-human primate, Macaca fascicularis. Scientific Reports, 9: 8504.
- Chew PC, Christianus A, Jaapar MZ, Ina-Salwany IS, Chong CM, Tan SG (2021) Microsatellite characterization of Malaysian mahseer (Tor spp.) for improvement of broodstock management and utilization. Animals, 11: 2633.
- Chun TC, Ling CK, Hee LC, Choo CS, Barin J, Fong AW, Abdullah MP, Tan SG (2018) Genetic diversity and inbreeding level in Deli dura and AVROS advanced breeding materials in oil palm (Elaeis guineensis Jacq.) using microsatellite markers. Journal of Oil Palm Research, 30: 366–79.
- Ee-Uli J, Yong CSY, Tan SG (2017) A glimpse into the genomic outlook of the long-tailed macaque (Macaca fascicularis). Pertanika Journal of Tropical Agricultural Science, 40: 329–41.
- Ee-Uli J, Yong CSY, Yeap SK, Alitheen NB, Rovie-Ryan JJ, Mat Isa N, Tan SG (2018) RNA sequencing of kidney and liver transcriptome obtained from wild cynomolgus macaque (Macaca fascicularis) originating from Peninsular Malaysia. BMC Research Notes, 11: 923.
- Ee-Uli J, Yong CSY, Yeap SK, Rovie-Ryan JJ, Mat Isa N, Tan SG, Alitheen NB (2017) RNA sequencing (RNA-Seq) of lymph node, spleen, and thymus transcriptome from wild Peninsular Malaysian cynomolgus macaque (Macaca fascicularis). PeerJ, 2017: e3566.
- Elsevier (2025) Scopus Content. www.elsevier.com/products/scopus/content (Assessed on 11 December 2025).
- Kristeen-Teo YW, Yeap SK, Tan SW, Omar AR, Aini I, Tan SG, Alitheen NB (2017) The effects of different velogenic NDV infections on the chicken bursa of Fabricius. BMC Veterinary Research, 13: 151.
- Kueh CL, Yong CY, Masoomi Dezfooli S, Bhassu S, Tan SG, Tan WS (2017) Virus-like particle of Macrobrachium rosenbergii nodavirus produced in Spodoptera frugiperda (Sf9) cells is distinctive from that produced in Escherichia coli. Biotechnology Progress, 33: 549–57.
- Ng WL, Yeap SK, Bakar NSMA, Jaafar WNWM, Tan SG (2016) Multiplex PCR assays for species discrimination of Cymbopogon citratus (DC.) Stapf and C. nardus (L.) Rendle, two common “serai” (lemon grass) species in Peninsular Malaysia. Sains Malaysiana, 45: 323–7.
- Noraini I, Tan SG, Gan YY, Teng YS (1980) Salivary peroxidase, Pm, and Ph protein polymorphisms in Malaysians. Human Genetics, 56: 205–7.
- Norakmal I, Tan SG (1979) Cerumen polymorphism in the three major ethnic groups of Malaysia. Japanese Journal of Human Genetics, 24: 119–21.
- Ong GH, Yap CK, Mahmood M, Tan SG, Hamzah MS (2017) Magnesium in local edible ulam (Centella asiatica) and its relation to their habitat soils in Peninsular Malaysia. Pertanika Journal of Tropical Agricultural Science, 40: 1–17.
- Pauzi AZM, Yeap SK, Abu N, Lim KL, Omar AR, Abd-Aziz SA, Chow ALT, Subramani T, Tan SG, Alitheen NB (2016) Combination of cisplatin and bromelain exerts synergistic cytotoxic effects against breast cancer cell line MDA-MB-231 in vitro. Chinese Medicine, 11: 46.
- Sarifudin MSA, Aminur Rahman MA, Yusoff FM, Arshad A, Tan SG (2017) Influence of salinity variations on the embryonic and early larval development of long-spined black sea urchin (Diadema setosum). Journal of Animal and Plant Sciences, 27: 316–24.
- Sarifudin MSA, Aminur Rahman MA, Yusoff FM, Arshad A, Tan SG (2016) Effects of temperature on the embryonic and early larval development in tropical species of black sea urchin, Diadema setosum (Leske, 1778). Journal of Environmental Biology, 37: 657–68.
- Seng TY, Ritter E, Mohamed Saad SH, Leao LJ, Singh RSH, Qamaruz-Zaman F, et al. (2016) QTLs for oil yield components in an elite oil palm (Elaeis guineensis) cross. Euphytica, 212: 399–425.
- Seyedi SS, Tan SG, Namasivayam P, Yong CSY (2016) Isolation and characterization of full-length cellulose synthase gene (HsCesA1) from roselle (Hibiscus sabdariffa L. var. UMKL). Sains Malaysiana, 45: 717–27.
- Siew GY, Ng WL, Salleh MF, Tan SW, Ky H, et al. (2018) Assessment of the genetic variation of Malaysian durian varieties using inter-simple sequence repeat markers and chloroplast DNA sequences. Pertanika Journal of Tropical Agricultural Science, 41: 321–31.
- Siew GY, Ng WL, Tan SW, Alitheen NB, Tan SG, Yeap SK (2018) Genetic variation and DNA fingerprinting of durian types in Malaysia using simple sequence repeat (SSR) markers. PeerJ, 2018: e4266.
- Tan SG (1976) Human saliva esterases: Genetic studies. Human Heredity, 26: 207–16.
- Tan SG, Ashton GC (1976) An autosomal glucose 6 phosphate dehydrogenase (hexose 6 phosphate dehydrogenase) polymorphism in human saliva. Human Heredity, 26: 113–23.
- Tan SG, Ashton GC (1976) Saliva acid phosphatases: Genetic studies. Human Heredity, 26: 81–9.
- Tan SG, Teng YS (1978) Saliva acid phosphatases and amylase in Senoi and Aboriginal Malays and superoxide dismutase in various racial groups of Peninsular Malaysia. Japanese Journal of Human Genetics, 23: 133–8.
- Tan SG, Teng YS (1979) Human saliva as a source of biochemical genetic markers: I. Techniques. Human Heredity, 29: 69–76.
- Tan SG, Teng YS (1979) Saliva acid phosphatases in Malaysians: Report of a new variant. Human Heredity, 29: 61–3.
- Tan SG, Gan YY, Asuan K, Abdullah F (1981) Gc subtyping in Malaysians and in Indonesians from North Sumatra. Human Genetics, 59: 75–6.
- Tan SG, Teng YS, Ganesan J, Lau KY, Lie-Injo LE (1979) Biochemical genetic markers in the Kadazans of Sabah, Malaysia. Human Genetics, 49: 349–53.
- Teng YS, Tan SG (1979) Acid α-glucosidase in Malaysians: Population studies and the occurrence of a new variant. Human Heredity, 29: 2–4.
- Teng YS, Tan SG (1979) Genetic evidence of gene flow from Indians to Malays. Japanese Journal of Human Genetics, 24: 1–8.
- Teng YS, Tan SG (1979) Human saliva as a source of biochemical genetic markers: II. Genetic interpretations and possible utilization. Human Heredity, 29: 129–33.
- Teng YS, Tan SG, Lopez CG, Ng T, Lie-Injo LE (1978) Genetic markers in Malaysians: Variants of soluble and mitochondrial glutamic oxaloacetic transaminase and salivary and pancreatic amylase, phosphoglucomutase III and saliva esterase polymorphisms. Human Genetics, 41: 347–54.
- Teng YS, Tan SG, Ng T, Lopez CG (1978) Red cell glyoxalase I and placental soluble aconitase polymorphisms in the three major ethnic groups of Malaysia. Japanese Journal of Human Genetics, 23: 211–5.
- Valdiani A, Talei D, Lattoo SK, Ortiz R, Rasmussen SK, Batley J, et al. (2017) Genoproteomics-assisted improvement of Andrographis paniculata: Toward a promising molecular and conventional breeding platform for autogamous plants affecting the pharmaceutical industry. Critical Reviews in Biotechnology, 37: 803–16.
- Van Eck NJ, Waltman L (2010) Software survey: VOSviewer, a computer program for bibliometric mapping. Scientometrics, 84: 523–38.
- Van Eck NJ, Waltman L (2014) Visualizing bibliometric networks. In Y. Ding, R. Rousseau, & D. Wolfram (Eds.), Measuring scholarly impact: Methods and practice, 285–320.
- Van Eck NJ, Waltman L (2017) Citation-based clustering of publications using CitNetExplorer and VOSviewer. Scientometrics, 111: 1053–70.
- Yee EYS, Zainuddin ZZ, Ismail A, Yap CK, Tan SG (2016) Molecular sex identification of painted storks (Mycteria leucocephala): Using FTA cards, horizontal PAGE and quicksilver staining. Journal of Genetics, 93: 15–8.
Article Information
Review Article
Received Date: December 20, 2025
Accepted Date: January 04, 2026
Published Date: January 08, 2026
Honouring a Scientific Pioneer: The Enduring Legacy of Professor Soon Guan Tan (1947–2023) in Malaysian Genetics, Biodiversity, and Environmental Science
Volume 5 | Issue 1
Citation
Chee Kong Yap, Wan Hee Cheng, Wen Siang Tan, Weiyun Chew, Franklin Berandah Edward (2025) Honouring a Scientific Pioneer: The Enduring Legacy of Professor Soon Guan Tan (1947–2023) in Malaysian Genetics, Biodiversity, and Environmental Science. J Environ Sci Energy 5: 1-10
Copyright
©2026 Chee Kong Yap. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.

doi: jese.2026.5.101

